Introduction to Porphyromonas gingivalis (P. gingivalis)
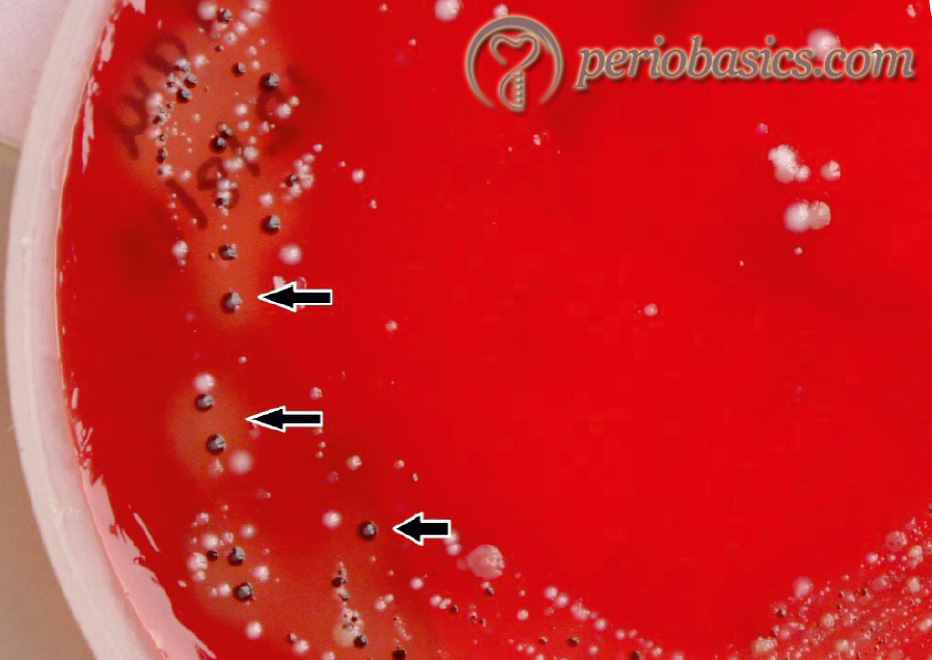
P. gingivalis is a Gram-negative, anaerobic, non-motile, asaccharolytic and black pigmented rod that form greenish-black colonies on blood agar plates 137. It is one of the major pathogens of chronic periodontitis 138, 139. This microorganism has been included in the red complex, which is strongly associated with periodontal destruction. In both periodontitis and healthy subjects, P. gingivalis can be recovered in low frequency from the subgingival flora, tongue, buccal mucosa, tonsils and saliva 140-142.
Structural components
Capsule
It is an anti-phagocytosis virulence factor. Six distinct capsular serotypes have been described (K1-K6) 143, 144. Capsules from three P. gingivalis strains have been purified. They are mainly composed of sugars like galactose, glucose, glucosamine, rhamnose, and mannose. A strong relationship exists between the extent P. gingivalis encapsulation and its ability to act as a pathogen. Increased encapsulation is related to decreased auto-agglutination, increased resistance to phagocytosis, serum resistance and decreased induction of PMN leukocyte chemiluminescence. Studies in a mouse infection model have revealed that encapsulated P. gingivalis strains are more virulent than non-encapsulated strains 58, 145-147. The components of the bacterial capsules have been used to make the vaccine.
Outer membrane
The outer membrane of P. gingivalis contains at least 20 major proteins, ranging from 20-90 kDa. These proteins have a significant effect on fibroblasts. They stimulate their proliferation. Because of this, a 24 kDa protein has been given name “fibroblast activating factor”. P. gingivalis also produces a 75 kDa major outer membrane protein that exists as a high molecular weight polymer. It has been found that these proteins can stimulate polyclonal B-cell invasion and can elicit IL-1 production. These outer membrane proteins are also important in plaque formation and coaggregation. P. gingivalis plays an important role in the formation and maintenance of plaque biofilm. Gibbson and Nygaard (1970) 57 demonstrated that …….. Contents available in the book………. Contents available in the book…… Contents available in the book…… Contents available in the book..
Periobasics: A Textbook of Periodontics and Implantology
The book is usually delivered within one week anywhere in India and within three weeks anywhere throughout the world.
India Users:
International Users:
Lipopolysaccharide (LPS)
The analysis of LPS from 6 different strains of P. gingivalis indicate that it contains sugars like rhamnose, mannose, and galactose. LPS has been studied for various immunological properties. Chemically, it is composed of 3 parts O-antigen, core, and lipid A. Endotoxin activity is confined to lipid A, while significant immunological activities are due to O antigen. LPS also interacts with CD14 receptors on cells and initiate the immune reaction.
Bacterial fimbriae
Fimbriae or pili are proteinaceous, filamentous appendages that protrude outwards from the bacterial cell surface and play a crucial role in virulence by stimulating bacterial attachment to host cells or tissues 148. Fimbriae of P. gingivalis were first recognized on the outer surface by electron microscopic observation 149, 150. They are arranged in a peritrichous pattern on P. gingivalis. Fimbrae is composed of 1000 units of fimbrillin. P. gingivalis fimbriae possess a strong ability to interact with host proteins (such as salivary proteins, extracellular matrix proteins), epithelial cells, and fibroblast, which promote the colonization of P. gingivalis in the oral cavity 151, 152. Animal experiments conducted strongly implicate fimbriae as an important virulence factor. Environmental factors like temperature, pH, haemin limitation, serum, saliva, osmotic effects and effects of Ca++ limitation have a significant effect on fimbrae formation. P. gingivalis fimbriae are classified into six genotypes based on the diversity of the fimA genes encoding each fimbrillin (types I to V, and type Ib), and that P. gingivalis with type II fimA is most closely associated with the progression of chronic periodontitis 153-155.
Proteinases
P. gingivalis produces a large number of hydrolytic, proteolytic and lipolytic enzymes that are produced essentially by all of the known strains. Many of these enzymes are exposed on the surface of the bacterium where they come in contact with the host cells and tissues. These enzymes play a significant role in periodontal disease progression, including dissemination of P. gingivalis into deeper tissues. Classification of proteinases is relied upon their catalytic functions. To date, four proteinases have been recognized: serine, aspartate, thiol, and metalloproteinases. The arg- and lys- proteinases have been given a common name Gingipains. There are at least 3 different genes which encode for the proteolytic activity of P. gingivalis. These genes encode for 2 cystein proteinases, arginine- gingipain (Arg-gingipain A & B; Rgp-A and Rgp-B) and lysine-gingipain (lys-gingipain, Kgp). Arg-gingipain is encoded by two genes and lys- gingipain by one gene 156. Movement of cysteine proteinases from the bacterial cytoplasm to the membrane involves the secretory pathway. Arg- and lys- proteinases or the gingipains belong to trypsin-like proteinases.
Colonization of P. gingivalis
Despite the host defense mechanisms in saliva and GCF, P. gingivalis can adhere and then colonize in gingival crevices to a variety of surface components lining the gingival crevicular cells and the tooth surface. The adhesion is mainly mediated by the fimbriae, although other bacterial components such as vesicles, hemagglutinin, and proteases may play an adjunctive role 157. P. gingivalis is capable of co-aggregating with Actinomyces naeslundii genospecies 2 (Actinomyces viscosus), Streptococcus gordonii, and Streptococcus mitis 158, 159.
Virulence factors associated with P. gingivalis
There are various virulence factors that have been demonstrated to be the key elements in the ability of P. gingivalis to evade the host defense and cause periodontal disease progression. These virulence factors include the following
- Fimbriae
- Proteinases
- Gingipains
- Collagenases
- Gelatinases
- Dipeptidyl aminopeptidase IV
- Hemagglutinin
- Capsule lipopolysaccharide (LPS)
- Outer membrane proteins (OMPs)
- Outer membrane vesicles (OMV’s)
Pathogenic process of P. gingivalis
The initial event in the pathogenicity of P. gingivalis is its interaction (adherence) in the oral cavity 160, 161. This process is facilitated by fimbriae, proteases, hemagglutinins and lipopolysaccharide 158. P. gingivalis fimbriae help in the bacterial interactions with the host cells and also with other bacteria. Bacterial adherence to mucosal and tooth surfaces, as well as bacterial co-aggregation, are the essential steps for colonization of various oral bacterial species. Fimbriae are involved in each of these processes 163, as well as cell adhesion molecules, including intercellular adhesion molecule 1 (ICAM-1), vascular cell adhesion molecule 1 (VCAM-1), and P- and E-selectins. P. gingivalis major fimbriae have been shown to be necessary for bacterial invasion in the host cells 163.
Proteinases
P. gingivalis proteinases have been reported to exhibit enzymatic activity against a broad range of host proteins, including host proteinase inhibitors, immunoglobulins, iron transporting/sequestering proteins, extracellular matrix proteins, bactericidal proteins and peptides, and proteins involved in the coagulation, complement, and kallikrein/kinin cascades 164. The majority of this activity is due to cysteine proteinases, referred to as gingipains.
Gingipains
Gingipains were originally considered as “trypsin-like proteases”, but actually, they comprise a group of cysteine endopeptidases 156 that have been reported to account for at least 85% of the general proteolytic activity displayed by P. gingivalis and 100% of the expressed “trypsin-like activity”. They are either secreted or membrane-bound and are arginine or lysine-specific. Gingipains are present in large quantities on the cell surface of P. gingivalis, and they can significantly contribute to the virulence exhibited by this species. They help in binding of the bacterium to the host tissues. They are classified into two groups based on substrate specificity. Gingipains-R (RgpA and RgpB) cleaves proteins after arginine residue and are encoded by two similar genes, RgpA and RgpB, while gingipains-K (Kgp) cleaves proteins after lysine residue. RgpA together with RgpB, accounts for all the trypsin-like activity of P. gingivalis. Kgp plays an important role in iron acquisition by binding to hemoglobin 156. They have been shown to have …….. Contents available in the book………. Contents available in the book…… Contents available in the book…… Contents available in the book..
Structural description of gingipains:
The mature form of RgpA possesses both, catalytic and hemagglutinin domain, while RgpB possesses only a catalytic domain. Hemagglutinin domain is responsible for the adherence of the microorganism to erythrocytes. There is a high degree of homology between the catalytic domains of RgpA and RgpB at both the DNA and protein levels, while the hemagglutinin domain of RgpA is similar to the P. gingivalis hemagglutinin domain of Kgp.
Mechanism of action of Gingipains
Vascular permeability enhancement:
R-gingipains are very potent factors of vascular permeability enhancement. This activity is induced through plasma prekallikrein activation and subsequent bradykinin release 166, 167. Gingipain K by itself is not able to induce vascular permeability in human plasma, but working in association with R gingipains, it can induce vascular permeability by cleaving bradykinin directly from high molecular weight kininogen 168. So, gingipains are responsible for the increase in gingival crevicular fluid production at periodontitis sites infected with P. gingivalis.
Cleavage of complement components:
RgpA is a very efficient enzyme in the generation of C5a through direct cleavage of C5. C5a is a potent chemotactic factor that likely contributes to the significant leukocyte infiltration at P. gingivalis induced periodontitis lesions. It also degrades C3 and in this way eliminates the creation of C3-derived opsonins, thus rendering P. gingivalis resistant to phagocytosis 168, 169. This ultimately results in massive accumulation of neutrophils in the inflamed periodontal tissue which contributes to very high levels of active granular proteinases (elastase, cathepsin G, gelatinase, and collagenases) in gingival crevicular fluid 170, 171 that may be responsible for connective tissue destruction.
Change in the subgingival environment:
The massive accumulation of neutrophils leads to the generation of high level of active granular proteinases which enables subgingival plaque bacteria to thrive due to the presence of high concentrations of peptides and amino acids, thus further aggravating tissue destruction 172.
Degradation of clotting factors:
Recent studies indicate that RgpA is capable of activating factor X and suggest that this gingipain could be responsible for the production of thrombin 173. Fibrinogen is a major target for Kgp. In vitro, this enzyme degrades the fibrinogen A alpha-chain within a minute 174, thus rendering it nonclottable. It has been postulated that the nonrestricted activity of gingipain K in periodontal pockets contributes to bleeding tendency, especially because it also very efficiently destroys the pro-coagulant portion of high-molecular-weight kininogen 174.
Stimulation of different cell types to produce inflammatory mediators:
More recently, it has been reported that P. gingivalis gingipains can activate different cell types leading to the secretion of inflammatory mediators 175-177. In one recent study, it has been found that P. gingivalis gingipains induced an inflammatory response in macrophages through the activation of intracellular kinases. Along with that, it was shown that both the Arg- and Lys-gingipain preparations induced the production of TNF-α and IL-8 by macrophages 178.
Collagenase:
The collagenases produced by P. gingivalis are capable of degrading collagen Type I. Proteolytic enzymes like gingipains, collagenases and hyaluronidases destroy periodontal tissue directly or indirectly, leading to the progression of the disease 158, 179. Numerous studies have demonstrated the collagenase activity of P. gingivalis 180-183. It has been shown that inactivation of gingipain R (both RgpA and RgpB) completely eliminates the capacity of P. gingivalis to cleave native collagen Type I, suggesting that …….. Contents available in the book………. Contents available in the book…… Contents available in the book…… Contents available in the book..
Gelatinase:
There is a scanty literature regarding gelatinases produced by P. gingivalis. However, they are considered as important components of total proteolytic activity demonstrated by this microorganism.
Dipeptidyl aminopeptidase IV:
It is a serine protease that cleaves X-Pro or X-Ala dipeptide at the N-terminal end of the polypeptide chain 185. It doesn’t have gelatinase activity itself, but has exopeptidase activity 186. These proteases along with other proteases produced by P. gingivalis degrade connective tissue, facilitating the invasion by P. gingivalis.
Hemagglutinin
P. gingivalis RgpA and Kgp have hemagglutinin domains. Their ability to hemagglutinate erythrocytes has been well documented 187, 188. Proteinase-hemagglutinin complexes may thus be important in the uptake of essential growth factors by the bacteria, via hemagglutination 189.
As gingipains are responsible for evasion of the host response, thus, it indicates that inhibition of gingipains should be a useful tool, both to assess the contribution of their proteolytic activities to the virulence of the bacterium and to facilitate the development of new therapeutic approaches to periodontal diseases. Research has been done to find out therapeutic agents that inhibit gingipain activity. Among this series of compounds used for gingipain inhibition, it has been found that KYT-1 and KYT-36 had the most potent and selective inhibitory activities on Rgp and Kgp, respectively 190.
Capsule lipopolysaccharide (LPS)
P. gingivalis like other Gram-negative bacteria are sheathed by a capsular LPS. This LPS capsule is outer membrane component recognized by the host cell, thus initiating intracellular signaling events. The affinity of LPS to its pattern recognition receptors, such as the TLRs and CD14, facilitates its recognition and discrimination from other bacterial species by the host immune system. P. gingivalis LPS has been shown to stimulate pro-inflammatory cytokine production by mono-cytes in vitro 191-194. However, it is a weaker cytokine stimulator as compared to the LPS of other Gram-negative bacteria 195. Structurally, LPS of P. gingivalis is different from other Gram-negative bacteria in terms of O-antigen, acylation patterns and receptor-activating capacities of the lipid A component 196.
Know More…
Vaccination against P. gingivalis:
Many researchers have investigated a group of cell surface carbohydrates designated as K-antigens 199, lipopolysaccharides 200, 201 and various proteins, including fimbriae 202, the 53-KDa and 67K-Da cell surface proteins 203, hemagglutinin 204 and cysteine proteases referred to as gingipains 205, for making a vaccine against P. gingivalis. Gingipains are present in large quantities on the cell surface of P. gingivalis and have the highest potential to be used as a vaccine antigen.
Gingipains as a candidate for periodontal vaccine:
Gingipain vaccines are mainly DNA vaccines. Arg-gingipain-encoding genes have been cloned from various P. gingivalis strains 206, 207. The sequence analysis shows that two separate genes located on the chromosome of P. gingivalis encode Rgp (RgpA and RgpB). As different gingipains contain a catalytic domain and a haem-agglutinin domain or any one of them, studies have been done to find out the protective host response against these after immunization 208-211. The antibody response has been documented against these antigens. Further studies are required in this direction to analyze the effectiveness of the vaccine and its practical implications.
Outer membrane Proteins (OMP’s)
The cell envelope of Gram-negative bacteria such as Porphyromonas species comprises of two cell membranes, the outer membrane and the inner membrane. The outer membrane is involved in most of the specific recognition processes as it is the most exposed region of a bacterial cell. The most abundant outer membrane proteins consistently identified are porins and OmpA-like proteins.
Outer membrane vesicles (OMV’s)
P. gingivalis produces OMV’s that carry virulence factors to the cell surface. These vesicles are released in an extracellular manner, retaining the full components of outer membrane constituents, including lipopolysaccharide, muramic acid, capsule, fimbriae, and gingipains 197,198. These measure between 50 and 250 nm in diameter, but are usually around 50 nm. These vesicles are primarily involved in communication with the host cells and other members of microbial biofilms, resulting in the transmission of virulence factors into these host cells and the formation of bacterial communities in plaque biofilm.
Conclusion
P. gingivalis is an important keystone pathogen. It has been demonstrated that it has all the major virulence factors that have been found to be associated with the progression of periodontal diseases. Along with this, it is a major candidate for preparation of the periodontal vaccine. However, we are still trying to understand the exact nature of the bacteria and a lot of research is required to understand the exact activity of this bacteria in progression of periodontal disease.
References
References are available in the hard copy of the website.
Periobasics: A Textbook of Periodontics and Implantology
The book is usually delivered within one week anywhere in India and within three weeks anywhere throughout the world.